Evaluating rrna as an indicator of microbial activity in environmental communities: limitations and uses
- Select a language for the TTS:
- UK English Female
- UK English Male
- US English Female
- US English Male
- Australian Female
- Australian Male
- Language selected: (auto detect) - EN
Play all audios:

ABSTRACT Microbes exist in a range of metabolic states (for example, dormant, active and growing) and analysis of ribosomal RNA (rRNA) is frequently employed to identify the ‘active’
fraction of microbes in environmental samples. While rRNA analyses are no longer commonly used to quantify a population’s growth rate in mixed communities, due to rRNA concentration not
scaling linearly with growth rate uniformly across taxa, rRNA analyses are still frequently used toward the more conservative goal of identifying populations that are currently active in a
mixed community. Yet, evidence indicates that the general use of rRNA as a reliable indicator of metabolic state in microbial assemblages has serious limitations. This report highlights the
complex and often contradictory relationships between rRNA, growth and activity. Potential mechanisms for confounding rRNA patterns are discussed, including differences in life histories,
life strategies and non-growth activities. Ways in which rRNA data can be used for useful characterization of microbial assemblages are presented, along with questions to be addressed in
future studies. SIMILAR CONTENT BEING VIEWED BY OTHERS PRIORITY EFFECTS IN MICROBIOME ASSEMBLY Article 27 August 2021 MICROBIOME DIFFERENTIAL ABUNDANCE METHODS PRODUCE DIFFERENT RESULTS
ACROSS 38 DATASETS Article Open access 17 January 2022 MIDAS 4: A GLOBAL CATALOGUE OF FULL-LENGTH 16S RRNA GENE SEQUENCES AND TAXONOMY FOR STUDIES OF BACTERIAL COMMUNITIES IN WASTEWATER
TREATMENT PLANTS Article Open access 07 April 2022 INTRODUCTION Microorganisms have essential roles in shaping and controlling virtually all ecosystems including the atmosphere, oceans,
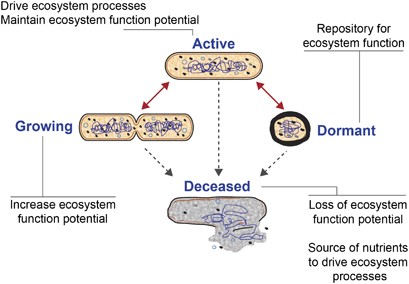
soils and plant- and animal-associated biomes. Microbes exist in different metabolic states in these systems: _growing_, _active_, _dormant_ and recently _deceased_ (Figure 1). These
metabolic states correspond to different degrees of influence that microbes can have on their environment. Therefore, to understand the relationships between microbial community structure
and ecosystem functions, it is important to accurately associate microbial identity with concurrent metabolic state. Simultaneous identification of microbes and their metabolic states has
been a longstanding goal in microbial ecology, and methods to achieve this have recently been accumulating in our molecular toolboxes. Nucleic-acid analysis has proven to be effective for
characterizing the phylogenetic, taxonomic and functional structure of microbial assemblages, but this approach has limitations when attempting to assess current metabolic state. Ribosomal
RNA genes (rRNA genes) are frequently used to identify microorganisms present in environmental samples regardless of metabolic state, while ribosomal RNA (rRNA) has been widely applied to
characterize the growing or active microbes. We found >100 studies that used rRNA for these purposes, including recent studies using rRNA to identify currently active microbes (for
example, Muttray and Mohn, 2000; Duineveld et al., 2001; Mills et al., 2005; Schippers et al., 2005; Gentile et al., 2006; DeAngelis et al., 2010; Jones and Lennon, 2010; Brettar et al.,
2011; Egert et al., 2011; Gaidos et al., 2011; Wüst et al., 2011; Mannisto et al., 2012; Hunt et al., 2013). However, conflicting patterns between rRNA content and growth rate indicate that
rRNA is not a reliable metric for growth or activity and in some cases may be grossly misleading. Virtually all molecular characterization methods are imperfect, but we suggest that using
rRNA analyses to evaluate microbial assemblages requires that limitations and underlying assumptions be clearly identified and understood. Here, we explore critical limitations and potential
causes of inconsistent rRNA/activity relationships. We then suggest employing rRNA abundance data as an index of potential activity and propose a framework for future application. The
reader should note that RNA extraction methods are important in interpreting the validity of any downstream RNA-based results. Often in the literature, purification and analytical methods
for RNA differ and are not shown to be reproducible and quantitative. As techniques advance, methods are continuously improved and new experimental results are presented. From a technical
point of view, it is extremely arduous to re-interpret older results based on new methodological improvements and is beyond the scope of this review. However, from an epistemological point
of view, it is important to keep in mind potential methodological biases to ensure that the assumptions of the relationship between RNA and activity are clearly articulated, and to recognize
the specific limitations of applying a broad generalization for RNA content to environmental samples. With this in mind, we discuss studies that utilized several different experimental
approaches; thus, observed discrepancies between rRNA abundance and activity are very likely to be at least in part biological in origin and not simply methodological artifact. We focus on
bacteria, which have been extensively studied, but many of the limitations discussed here are likely relevant for other microbes, including archaea, fungi and algae. RRNA AND ITS USE IN
MICROBIAL ECOLOGY The cell’s total RNA pool is mainly composed of rRNA (82–90%) (Tissieres and Watson, 1958; Neidhardt and Magasanik, 1960; Neidhardt, 1987). As an integral structural
component of ribosomes, rRNA is a fundamental constituent of all known microorganisms and most rRNA found in a cell is ribosome associated (Lindahl, 1975; Nomura et al., 1984). Total RNA
concentration is generally proportional to rRNA concentration and to the number of ribosomes in the cell, and has often been employed as a proxy for both (Kerkhof and Ward, 1993; Poulsen et
al., 1993; Bremer and Dennis, 1996). In pure-culture experiments, cell counts can be done to determine RNA or ribosome concentration per cell. In mixed communities, other methods of
normalization are necessary. Commonly, RNA or rRNA concentration is normalized to the number of cells using DNA concentration to calculate the RNA:DNA or an rRNA:rRNA gene ratio (for
example, Kemp et al., 1993; Kerkhof and Ward, 1993; Poulsen et al., 1993; Muttray et al., 2001), since DNA concentration per cell is generally more stable than RNA concentration. Note,
however, that while cell genome content commonly varies less than RNA content, genome abundance per cell can vary significantly and therefore could influence RNA:DNA measurements (Schaechter
et al., 1958; Cooper and Helmstetter, 1968; Sukenik et al., 2012), but this issue will not be addressed here. Historically, rRNA analyses have been used to quantify populations’ growth
rates in mixed microbial communities (for example, Poulsen et al., 1993; Muttray et al., 2001), but recent application has shifted toward the more qualitative approach using rRNA to identify
currently active microbial populations in a mixed community (for example, Jones and Lennon, 2010; Kamke et al., 2010; Campbell et al., 2011; DeAngelis et al., 2011; Gaidos et al., 2011;
Reid et al., 2011; Baldrian et al., 2012; Mannisto et al., 2012; Mattila et al., 2012; Simister et al., 2012; Campbell and Kirchman, 2013; Hunt et al., 2013; Yarwood et al., 2013). Two
principal lines of evidence used to support rRNA as an indicator of current activity originate from earlier studies testing how rRNA scales with growth rate. First, total RNA and rRNA
content correlate well with growth rate for a handful of microbes in pure culture, over a wide range of growth rates under balanced growth conditions (that is, growing in an unchanging
environment) (Schaechter et al., 1958; Neidhardt and Magasanik, 1960; Rosset et al., 1966; Koch, 1970; Kemp et al., 1993; Kerkhof and Ward, 1993; Poulsen et al., 1993; Wagner, 1994; Bremer
and Dennis, 1996; Ramos et al., 2000). Second, decreased rRNA content is associated with decreased growth rate for some organisms growing under specific nutrient-limiting conditions
(Mandelstam and Halvorson, 1960; Davis et al., 1986; Kramer and Singleton, 1992; Tolker-Nielsen et al., 1997). Note that the relationship between rRNA concentration and growth rate is
frequently coupled with the assumption that activity and growth are synonymous. Here, we distinguish growth from activity; while all growing organisms are active, not all active organisms
are growing (Figure 1). Experimental evidence demonstrates numerous limitations to use rRNA to quantify population growth rates in mixed communities, many of which have been addressed in
methodological reviews (for example, Molin and Givskov, 1999). However, while most of these limitations are also pertinent when attempting to identify which microbes are currently active in
a community, these limitations are frequently overlooked or ignored in practice. Here, we provide a summary of limitations (Box 1) that pertain to the relationship between rRNA and current
activity, and discuss relevant examples to assess the information that rRNA data can actually provide. BOX 1: LIMITATIONS OF RRNA AS AN INDICATOR OF CURRENT MICROBIAL ACTIVITY (REFERENCES
INCLUDE THE SEMINAL STUDIES THAT WERE LATER OFTEN OVERLY GENERALIZED TO SUPPORT RRNA–ACTIVITY RELATIONSHIP) * 1 Concentration of rRNA and growth rate are not always simply correlated;
therefore, the relationship between rRNA and activity is not likely consistent (Schaechter et al., 1958; Mandelstam and Halvorson, 1960; Flärdh et al., 1992, Kemp et al., 1993;
Tolker-Nielsen et al., 1997; Binder and Liu, 1998; Lepp and Schmidt, 1998; McKillip et al., 1998; Kerkhof and Kemp, 1999; Morgenroth et al., 2000; Oda et al., 2000; Schmid et al., 2001;
Worden and Binder, 2003). * 2 The relationship between rRNA concentration and growth rate can differ significantly among taxa; therefore, relative rRNA abundance will likely not provide
robust information regarding which taxa are relatively more active in a community (Mandelstam and Halvorson, 1960; Wade and Robinson, 1965; Rosset et al., 1966; Kemp et al., 1993; Pang and
Winkler, 1994; Oda et al., 2000; Binnerup et al., 2001; Worden and Binder, 2003). * 3 Dormant cells can contain high numbers of ribosomes; therefore, in environments that could likely
contain dormant cells, employing rRNA to identify current activity is highly problematic (Chaloupecky, 1964; Bishop and Doi, 1966; Chambon et al., 1968; Filion et al., 2009; Sukenik et al.,
2012). * 4 The relationship between non-growth activities and concentration of rRNA has not yet been investigated. CRITICAL ANALYSIS OF RRNA AS AN INDICATOR OF CURRENT ACTIVITY CONCENTRATION
OF RRNA AND GROWTH RATE ARE NOT ALWAYS SIMPLY CORRELATED The first line of evidence that has been used to support a predictable relationship between the presence of rRNA and current
activity is based on pure-culture studies assessing growth under balanced growth conditions. However, even under constrained conditions (balanced growth) the correlation between growth rate
and rRNA concentration is commonly not straightforward and in some cases breaks down altogether. For example, the relationship between growth rate and rRNA content is not linear or
consistent across all measured growth rates. Under balanced growth conditions, _Synechococcus_ and _Prochlorococcus_ strains can have a three-phase relationship between growth and rRNA
concentration: (1) at low growth rates, rRNA concentration remains constant, (2) at intermediate growth rates, rRNA concentration increases linearly with growth rate and (3) at higher growth
rates, rRNA content decreases as growth rate increases (Binder and Liu, 1998; Worden and Binder, 2003). For these organisms, rRNA concentration is not a robust proxy for growth rate. We
argue that rRNA will also not be a robust measure of current activity, since changes in growth-associated activity must impact total activity. Additionally, balanced growth conditions are
unlikely in most environments. Little work has characterized how rRNA concentration varies with growth rate under more environmentally realistic non-steady state conditions. Kerkhof and Kemp
(1999) identified three different relationship patterns between rRNA concentration and growth rate for Proteobacteria strains under non-steady state conditions: a direct linear
relationship, an indirect relationship in which cell growth rate consistently lagged behind rRNA concentration or no discernible relationship. The latter was observed in _Vibrio fischeri_,
and included periods during which growth rate decreased while rRNA content increased. Again, since growth activity likely accounts for much of total activity, these results indicate that
using rRNA concentration to assess current activity or changes in activity over time is problematic. Further evidence showing potential for misleading environmental interpretations includes
significant increase in cellular rRNA in _Aphanizomenon ovalisporum_ cells transitioning from vegetative to dormant state (Sukenik et al., 2012). These results indicate that a measurable
increase in rRNA abundance does not necessarily indicate an increase in activity. A second line of evidence cited to support rRNA as an indicator of current activity arises from studies on
RNA stability under different growth-limiting conditions (for example, carbon or nutrient limitations). Several studies have reported that exponentially growing cells subjected to nutrient
starvation degrade much of their rRNA in a relatively short time. However, the dynamics of cellular rRNA may be strongly tied to previous growth conditions. For example, _Azotobacter agilis_
was grown on different substrates, then starved for 72 h (Sobek et al., 1966). When grown on glucose, RNA did not decrease during the starvation period, but O2 consumption dramatically
dropped, indicating that cell activity (that is, respiration) declined. In another study, _Rhodopseudomonas palustris_ cells were grown at different growth rates, then carbon-starved (Oda et
al., 2000). The rRNA concentration of _R. palustris_ cells grown at maximum growth rate decreased by ∼50% within a week of starvation; however, cells grown at lower rates before the
starvation period were able to maintain near pre-starvation rRNA concentrations for more than a week of starvation. These results indicate that measurable rRNA concentration can be
influenced not only by current conditions, but also by life history (that is, the sequence of events that impacted an organism up to a given time point, and the resulting physiological
response to these events). THE RELATIONSHIP BETWEEN RRNA CONCENTRATION AND GROWTH RATE CAN DIFFER SIGNIFICANTLY AMONG TAXA Relating rRNA concentration and growth rate becomes even more
problematic when considering microbial assemblages. rRNA concentration may correlate well with growth rate in some strains of bacteria, but correlations can differ significantly between
strains (Wade and Robinson, 1965; Kemp et al., 1993; Pang and Winkler, 1994; Binnerup et al., 2001; Worden and Binder, 2003). Even at the ‘species’ level of bacteria, the relationship
between rRNA and growth rate can differ significantly between subpopulations (Rosset et al., 1966; Licht et al., 1999). Hence, using rRNA to compare relative activity or changes in activity
between taxa will likely provide misleading information. DORMANT CELLS CAN CONTAIN HIGH NUMBERS OF RIBOSOMES Dormant organisms contain measurable amounts of rRNA (Chambon et al., 1968) and
in some cases can contain significantly more rRNA in dormancy than in a vegetative state (Sukenik et al., 2012). Detectability of rRNA in dormant cells can be affected more by methodology
(due to changes in cell structure) than by low levels of rRNA (Filion et al., 2009). The issue of dormant cells containing measurable rRNA concentrations can be especially problematic when
using rRNA data to identify currently active organisms in environments likely to contain many dormant organisms such as soil, deep subsurface, frozen environments or the atmosphere. One
approach to discounting rRNA in dormant cells is to estimate the rRNA concentration per cell for specific taxa by calculating rRNA:rRNA gene ratios, then defining a minimum cutoff value for
activity (for example, DeAngelis et al., 2011; Jones and Lennon, 2010). However, rRNA:rRNA gene ratios have been characterized for very few bacteria in dormant state. The limited available
evidence demonstrates the difficulties in establishing a suitable universal cutoff value for rRNA:rRNA gene ratio. For example, an RNA:DNA ratio of around 5 was found both in dormant
_Bacillus megaterium_ (Chambon et al., 1968) and in bacteria growing at the rapid pace of ∼0.5 h−1 (Kerkhof and Ward, 1993). THE RELATIONSHIP BETWEEN NON-GROWTH ACTIVITIES AND CONCENTRATION
OF RRNA HAS NOT BEEN INVESTIGATED Finally, the relationship between rRNA concentration and growth rate is commonly considered to be equivalent to that between rRNA and activity. However,
many microbial activities are not necessarily related to growth, including those associated with maintenance, such as cell motility, osmoregulation, defense against oxidative stress,
communication, exopolysaccharide production or conjugation (van Bodegom, 2007). To our knowledge, no published work has investigated the relationship between non-growth activities and rRNA
concentration. It has been hypothesized that under certain stress conditions, microbes can dramatically increase the portion of metabolism geared toward non-growth maintenance activities
(Schimel et al., 2007), indicating that, under appropriate conditions, non-growth activities may contribute significantly to ecosystem processes. RELATIONSHIP BETWEEN RRNA, GROWTH AND
ACTIVITY: PHYSIOLOGICAL LINKS The multi-level modulation and regulation of most cell functions may easily invalidate simple correlations between current metabolic state and rRNA abundance.
For example, the relationship between microbial activity and measurable rRNA can be influenced by heterogeneity of cell physiology within a population (Licht et al., 1999), changes in the
ratio of non-growth to growth-specific metabolic activity, life history (Oda et al., 2000), life strategy (Flärdh et al., 1992; Lepp and Schmidt, 1998; Mitchell et al., 2009; Sukenik et al.,
2012), sample heterogeneity, changing environmental conditions and of course fundamental enzyme kinetics (that is, substrate concentration). Additionally, the concentration of rRNA in a
cell at a given point in time is the net result of rRNA synthesis (that is, transcription) and degradation rates (Gausing, 1977), each of which may be under distinct controls. All of these
factors can affect the relationship between ribosome turnover and microbial activity at multiple levels (Figure 2) and should be considered when analyzing rRNA data from environmental
samples. RRNA ANALYSES IN COMMUNITY ECOLOGY rRNA-based measurements can provide meaningful insight into microbial community dynamics. rRNA directly relates to a population’s potential to
catalyze the specific function of protein synthesis, and can therefore document the relative expression of this function. rRNA-based measurements provide a specific piece of information in
the spectrum of molecular approaches (including metagenomics, metatranscriptomics, metaproteomics and community proteogenomics) that are increasingly applied to study microbial communities.
Metagenomic data provide information about the functional potential of a sample, without providing insight into current metabolic state. Metatranscriptomic data come one step closer to
current metabolic state, without providing direct evidence of translation or enzyme activity. Metaproteomic data come an additional step closer to current metabolic state, by identifying
enzymes expressed in a community, but without providing direct evidence of enzyme activity. While rRNA is a product of transcription, community rRNA data are more analogous to metaproteomic
than to metatranscriptomic (mRNA) data; rRNA is generally much more stable than mRNA (Snyder and Champness, 2007), and is not translated to protein but instead acts as a structural component
of housekeeping catalysts (ribosomes). Therefore, rRNA data can provide evidence of the relative expression of an enzyme, with the explicit function of protein synthesis, for different
populations in a community. Analogously, in metaproteomics, environmental proteins are characterized to provide information about specific enzymatic functions that are expressed (Wilmes and
Bond, 2004). Taking this analogy one step further, the community proteogenomics approach can be used to map the expressed function of a community (metaproteomic data) onto the available
template of potential functions (metagenomic data) (Verberkmoes et al., 2009), to provide valuable information about how environmental changes correspond to changes in community expression
in the context of community composition. Similarly, rRNA data can be mapped onto rRNA gene data to illuminate relative ribosomal expression of the total community. However, it is important
to recognize that while enzyme/protein data come closer than gene and transcript data to identifying real-time activity, the presence of an enzyme does not unequivocally denote current
activity for a given function, because many factors control enzymatic activity _in vivo_ (Nannipieri et al., 2002). Similarly, the presence of rRNA is indicative of protein synthesis
_potential_, not of _realized_ protein synthesis (Figure 2). The number of ribosomes present at a given time limits the maximum protein synthesis activity for a population, but does not
directly inform about realized protein synthesis activity. The distinction between actual activity and potential activity is critical when attempting to identify and characterize the
dynamics of organisms that drive ecosystem functions (Figure 1). APPLICATIONS IN MICROBIAL ECOLOGY: FUTURE DIRECTIONS What does measuring ‘protein synthesis potential’ tell us about
microbial populations? The relationship between the number of ribosomes and the ability to synthesize proteins links the quantity of rRNA in a population with its potential for growth and
acclimation (that is, to upregulate or change currently expressed metabolic functions). rRNA can represent potential future activity, in addition to reflecting historical activity and
conditions (as discussed above). For example, some microorganisms increase ribosome concentration as they enter a dormant state, a life strategy that provides them with higher protein
synthesis potential, and therefore potentially higher fitness, as they return to a vegetative state when environmental conditions improve (Sukenik et al., 2012). Similarly, non-dormant
populations maintaining ribosome levels above current protein synthesis demands likely have the ability to rapidly shift metabolic functions to adapt to changing conditions, thereby becoming
better competitors (Koch, 1971; Alton and Koch, 1974; Flärdh et al., 1992). Recognizing that rRNA concentration reflects past, current and future activities in addition to different life
strategies restricts its utility as a metric of real-time activity, but provides the basis for generating and testing important hypotheses. Several studies show that under repeated temporal
patterns of changing environmental conditions, microbes may develop an anticipatory life strategy, enduring one phase of the cycle while preparing for a more favorable phase that regularly
follows. Further, accumulating or maintaining rRNA during periods of low metabolic activity may confer a competitive advantage during a favorable phase of the cycle. In _Synechococcus_ sp.
incubated under light and dark diurnal cycles, rRNA content increased during dark periods compared with light periods; in contrast, growth occurred during the light periods and ceased during
the dark periods (Lepp and Schmidt, 1998). Similar results were found for a strain of _Prochlorococcus_ in which expression of ribosomal genes was higher during a dark cycle than during a
light cycle (Zinser et al., 2009). Further evidence for anticipatory behavior in bacteria was found in _E. coli_ manifesting a Pavlovian-type response to a primary stimulus by preemptively
modifying genetic expression for a secondary stimulus before it occurred (Tagkopoulos et al., 2008; Mitchell et al., 2009). Anticipatory strategies may also take place on a seasonal scale:
at the end of a summer dry-down period, Mediterranean soil communities showed almost no measurable microbial activity (based on CO2 production), yet total extractable bacterial 16S rRNA was
similar to that found after the microbes become activated by the first wet-up event (Placella et al., 2012), which could reflect anticipation for the upcoming annual rainy season (Barnard et
al., 2013). If anticipatory life strategies reflected in rRNA concentrations are common in microbial populations experiencing repeated cyclic patterns, then can this information be
meaningfully applied to predict future changes in ecosystem function? To utilize rRNA data to characterize microbial assemblages, we need to better our understanding of how these data relate
to environmental conditions and community interactions; this understanding could be furthered by several experimental approaches: * a) _Coupling direct measurements of metabolic activity to
rRNA data_. * b) _Explicitly testing the relationship between non-growth activities and rRNA concentrations_. * c) _Characterizing ribosome turnover under different environmental
conditions._ CONCLUSION A number of pure-culture studies have shown a correlation between growth rate and rRNA concentration. This relationship makes intuitive and biological sense, since
rRNA is a critical component of ribosomes, and ribosomes are necessary to synthesize protein. However, the correlation between real-time activity and rRNA in environmental samples is
inconsistent due to differences in life histories, life strategies and non-growth activities. Using rRNA analysis as a general indicator of currently active microbes in environmental samples
is not valid under many circumstances, and may actually hinder progress connecting microbial activities to ecosystem functions. Considering rRNA measurements as indicators of protein
synthesis potential provides microbial ecologists with a robust framework, facilitating a more prudent yet comprehensive understanding of the complex dynamics at play in microbial
communities. REFERENCES * Alton TH, Koch AL . (1974). Unused protein synthetic capacity of Escherichia coli grown in phosphate-limited chemostats. _J Mol Biol_ 86: 1–9. Article CAS PubMed
Google Scholar * Baldrian P, Kolarík M, Štursová M, Kopecky J, Valášková V, Vētrovsky T _et al_ (2012). Active and total microbial communities in forest soil are largely different and
highly stratified during decomposition. _ISME J_ 6: 248–258. Article CAS PubMed Google Scholar * Barnard RL, Osborne CA, Firestone MK . (2013). Responses of soil bacterial and fungal
communities to extreme desiccation and rewetting. _ISME J_ e-pub ahead of print 4 July 2013; doi: 10.1038/ismej.2013.104. Article CAS PubMed PubMed Central Google Scholar * Binder BJ,
Liu YC . (1998). Growth rate regulation of rRNA content of a marine Synechococcus (cyanobacterium) strain. _Appl Environ Microbiol_ 64: 3346–3351. CAS PubMed PubMed Central Google Scholar
* Binnerup S, Bloem J, Hansen B, Wolters W, Veninga M, Hansen M . (2001). Ribosomal RNA content in microcolony forming soil bacteria measured by quantitative 16S rRNA hybridization and
image analysis. _FEMS Microbiol Ecol_ 37: 231–237. Article CAS Google Scholar * Bishop H, Doi R . (1966). Isolation and characterization of ribosomes from Bacillus subtilis spores. _J
Bacteriol_ 91: 695–701. CAS PubMed PubMed Central Google Scholar * Bremer H, Dennis PP . (1996) _Modulation of Chemical Composition and Other Parameters of the Cell by Growth Rate_ 2nd
edn. ASM Press: Washington, DC. Google Scholar * Brettar I, Christen R, HöFle MG . (2011). Analysis of bacterial core communities in the central Baltic by comparative RNA–DNA-based
fingerprinting provides links to structure–function relationships. _ISME J_ 6: 195–212. Article PubMed PubMed Central Google Scholar * Campbell BJ, Yu L, Heidelberg JF, Kirchman DL .
(2011). Activity of abundant and rare bacteria in a coastal ocean. _Proc Natl Acad Sci USA_ 108: 12776–12781. Article CAS PubMed PubMed Central Google Scholar * Campbell BJ, Kirchman DL
. (2013). Bacterial diversity, community structure and potential growth rates along an estuarine salinity gradient. _ISME J_ 7: 210–220. Article CAS PubMed Google Scholar * Chaloupecky
V . (1964). Ribosomes in growing and non-growing bacterial cells. _Folia Microbiol_ 9: 232–237. Article Google Scholar * Chambon P, DuPraw EJ, Kornberg A . (1968). Biochemical studies of
bacterial sporulation and germination. _J Biol Chem_ 243: 5101–5109. CAS PubMed Google Scholar * Cooper S, Helmstetter CE . (1968). Chromosome replication and the division cycle of
Escherichia coli B/r. _J Mol Biol_ 31: 519–540. Article CAS PubMed Google Scholar * Davis BD, Luger SM, Tai PC . (1986). Role of ribosome degradation in the death of starved Escherichia
coli cells. _J Bacteriol_ 166: 439–445. Article CAS PubMed PubMed Central Google Scholar * DeAngelis KM, Silver WL, Thompson AW, Firestone MK . (2010). Microbial communities acclimate
to recurring changes in soil redox potential status. _Environ Microbiol_ 12: 3137–3149. Article CAS PubMed Google Scholar * DeAngelis KM, Wu CH, Beller HR, Brodie EL, Chakraborty R,
DeSantis TZ _et al_ (2011). PCR Amplification-independent methods for detection of microbial communities by the high-density microarray PhyloChip. _Appl Environ Microbiol_ 77: 6313–6322.
Article CAS PubMed PubMed Central Google Scholar * Duineveld BM, Kowalchuk GA, Keijzer A, Van Elsas JD, Van Veen JA . (2001). Analysis of bacterial communities in the rhizosphere of
chrysanthemum via denaturing gradient gel electrophoresis of PCR-amplified 16S rRNA as well as DNA fragments coding for 16S rRNA. _Appl Environ Microbiol_ 67: 172. Article CAS PubMed
PubMed Central Google Scholar * Egert M, Schmidt I, Höhne H-M, Lachnit T, Schmitz RA, Breves R . (2011). Ribosomal RNA-based profiling of bacteria in the axilla of healthy males suggests
right-left asymmetry in bacterial activity. _FEMS Microbiol Ecol_ 77: 146–153. Article CAS PubMed Google Scholar * Filion G, Laflamme C, Turgeon N, Ho J, Duchaine C . (2009).
Permeabilization and hybridization protocols for rapid detection of Bacillus spores using fluorescence in situ hybridization. _J Microbiol Methods_ 77: 29–36. Article CAS PubMed Google
Scholar * Flärdh K, Cohen P, Kjelleberg S . (1992). Ribosomes exist in large excess over the apparent demand for protein-synthesis during carbon starvation in marine Vibrio sp. strain CCUG
15956. _J Bacteriol_ 174: 6780–6788. Article PubMed PubMed Central Google Scholar * Gaidos E, Rusch A, Ilardo M . (2011). Ribosomal tag pyrosequencing of DNA and RNA from benthic coral
reef microbiota: community spatial structure, rare members and nitrogen-cycling guilds. _Environ Microbiol_ 13: 1138–1152. Article PubMed Google Scholar * Gausing K . (1977). Regulation
of ribosome production in Escherichia coli: synthesis and stability of ribosomal RNA and of ribosomal protein messenger RNA at different growth rates. _J Mol Biol_ 115: 335–354. Article CAS
PubMed Google Scholar * Gentile G, Giuliano L, D'auria G, Smedile F, Azzaro M, De Domenico M _et al_ (2006). Study of bacterial communities in Antarctic coastal waters by a
combination of 16S rRNA and 16S rDNA sequencing. _Environ Microbiol_ 8: 2150–2161. Article CAS PubMed Google Scholar * Hunt DE, Lin Y, Church MJ, Karl DM, Tringe SG, Izzo LK _et al_
(2013). Relationship between abundance and specific activity of bacterioplankton in open ocean surface waters. _Appl Environ Microbiol_ 79: 177–184. Article CAS PubMed PubMed Central
Google Scholar * Jones SE, Lennon JT . (2010). Dormancy contributes to the maintenance of microbial diversity. _Proc Natl Acad Sci USA_ 107: 5881–5886. Article CAS PubMed PubMed Central
Google Scholar * Kamke J, Taylor MW, Schmitt S . (2010). Activity profiles for marine sponge-associated bacteria obtained by 16S rRNA vs 16S rRNA gene comparisons. _ISME J_ 4: 498–508.
Article CAS PubMed Google Scholar * Kemp P, Lee S, LaRoche J . (1993). Estimating the growth rate of slowly growing marine bacteria from RNA content. _Appl Environ Microbiol_ 59: 2594.
CAS PubMed PubMed Central Google Scholar * Kerkhof L, Ward B . (1993). Comparison of nucleic acid hybridization and fluorometry for measurement of the relationship between RNA/DNA ratio
and growth rate in a marine bacterium. _Appl Environ Microbiol_ 59: 1303. CAS PubMed PubMed Central Google Scholar * Kerkhof L, Kemp P . (1999). Small ribosomal RNA content in marine
Proteobacteria during non-steady-state growth. _FEMS Microbiol Ecol_ 30: 253–260. Article CAS PubMed Google Scholar * Koch AL . (1970). Overall controls on the biosynthesis of ribosomes
in growing bacteria. _J Theor Biol_ 28: 203–231. Article CAS Google Scholar * Koch AL . (1971). The adaptive responses of Escherichia coli to a feast and famine existence. _Adv Microbiol
Physiol_ 6: 147–217. Article CAS Google Scholar * Kramer JG, Singleton FL . (1992). Variations in rRNA content of marine Vibrio spp. during starvation-survival and recovery. _Appl Environ
Microbiol_ 58: 201. CAS PubMed PubMed Central Google Scholar * Lepp P, Schmidt T . (1998). Nucleic acid content of Synechococcus spp. during growth in continuous light and light/dark
cycles. _Arch Microbiol_ 170: 201–207. Article CAS PubMed Google Scholar * Licht TR, Tolker-Nielsen T, Holmstrøm K, Krogfelt KA, Molin S . (1999). Inhibition of Escherichia coli
precursor-16S rRNA processing by mouse intestinal contents. _Environ Microbiol_ 1: 23–32. Article CAS PubMed Google Scholar * Lindahl L . (1975). Intermediates and time kinetics of the
in vivo assembly of Escherichia coli ribosomes. _J Mol Biol_ 92: 15–37. Article CAS PubMed Google Scholar * Mandelstam J, Halvorson H . (1960). Turnover of protein and nucleic acid in
soluble and ribosome fractions of non-growing Escherichia coli. _Biochim Biophys Acta_ 40: 43–49. Article CAS PubMed Google Scholar * Mannisto MK, Kurhela E, Tiirola M, Haggblom MM .
(2012). Acidobacteria dominate the active bacterial communities of Arctic tundra with widely divergent winter-time snow accumulation and soil temperatures. _FEMS Microbiol Ecol_ 84: 47–59.
Article PubMed Google Scholar * Mattila HR, Rios D, Walker-Sperling VE, Roeselers G, Newton ILG . (2012). Characterization of the active microbiotas associated with honey bees reveals
healthier and broader communities when colonies are genetically diverse. _PLoS One_ 7: e32962. Article CAS PubMed PubMed Central Google Scholar * McKillip JL, Jaykus LA, Drake M .
(1998). rRNA stability in heat-killed and UV-irradiated enterotoxigenic Staphylococcus aureus and Escherichia coli O157:H7. _Appl Environ Microbiol_ 64: 4264–4268. CAS PubMed PubMed
Central Google Scholar * Mills HJ, Martinez RJ, Story S, Sobecky PA . (2005). Characterization of microbial community structure in Gulf of Mexico gas hydrates: comparative analysis of
DNA-and RNA-derived clone libraries. _Appl Environ Microbiol_ 71: 3235. Article CAS PubMed PubMed Central Google Scholar * Mitchell A, Romano GH, Groisman B, Yona A, Dekel E, Kupiec M
_et al_ (2009). Adaptive prediction of environmental changes by microorganisms. _Nature_ 460: 220–224. Article CAS PubMed Google Scholar * Molin S, Givskov M . (1999). Application of
molecular tools for in situ monitoring of bacterial growth activity. _Environ Microbiol_ 1: 383–391. Article CAS PubMed Google Scholar * Morgenroth E, Obermayer A, Arnold E, Brühl A,
Wagner M, Wilderer P . (2000). Effect of long-term idle periods on the performance of sequencing batch reactors. _Water Sci Technol_ 41: 105–113. Article CAS Google Scholar * Muttray AF,
Mohn WW . (2000). Quantitation of the population size and metabolic activity of a resin acid degrading bacterium in activated sludge using slot-blot hybridization to measure the rRNA:rDNA
ratio. _Microb Ecol_ 38: 348–357. Article Google Scholar * Muttray AF, Yu Z, Mohn WW . (2001). Population dynamics and metabolic activity of Pseudomonas abietaniphila BKME-9 within pulp
mill wastewater microbial communities assayed by competitive PCR and RT-PCR. _FEMS Microbiol Ecol_ 38: 21–31. Article CAS Google Scholar * Nannipieri P, Kandeler E, Ruggiero P . (2002).
Enzyme activities and microbiological and biochemical processes in soil. In: Burns RG, Dick RP (eds) _Enzymes in the Environment: Activity, Ecology, and Applications_. Marcel Dekker, Inc.:
New York, NY, pp 1–33. Google Scholar * Neidhardt FC, Magasanik B . (1960). Studies on the role of ribonucleic acid in the growth of bacteria. _Biochim Biophys Acta_ 42: 99–116. Article
CAS PubMed Google Scholar * Neidhardt FC . (1987) _Chemical Composition of Escherichia coli_ VOL. 1. American Society for Microbiology: Washington, DC. Google Scholar * Nomura M, Gourse
R, Baughman G . (1984). Regulation of the synthesis of ribosomes and ribosomal components. _Annu Rev Biochem_ 53: 75–117. Article CAS PubMed Google Scholar * Oda Y, Slagman S-J, Meijer
WG, Forney LJ, Gottschal JC . (2000). Influence of growth rate and starvation on fluorescent in situ hybridization of Rhodopseudomonas palustris. _FEMS Microbiol Ecol_ 32: 205–213. Article
CAS Google Scholar * Pang H, Winkler HH . (1994). The concentrations of stable RNA and ribosomes in Rickettsia prowazekii. _Mol Microbiol_ 12: 115–120. Article CAS PubMed Google Scholar
* Placella SA, Brodie EL, Firestone MK . (2012). Rainfall-induced carbon dioxide pulses result from sequential resuscitation of phylogenetically clustered microbial groups. _Proc Natl Acad
Sci USA_ 109: 10931–10936. Article CAS PubMed PubMed Central Google Scholar * Poulsen L, Ballard G, Stahl D . (1993). Use of rRNA fluorescence in situ hybridization for measuring the
activity of single cells in young and established biofilms. _Appl Environ Microbiol_ 59: 1354. CAS PubMed PubMed Central Google Scholar * Ramos C, Molbak L, Molin S . (2000). Bacterial
activity in the rhizosphere analyzed at the single-cell level by monitoring ribosome contents and synthesis rates. _Appl Environ Microbiol_ 66: 801. Article CAS PubMed PubMed Central
Google Scholar * Reid NM, Addison SL, Macdonald LJ, Lloyd-Jones G . (2011). Biodiversity of active and inactive bacteria in the gut flora of wood-feeding huhu beetle larvae (Prionoplus
reticularis). _Appl Environ Microbiol_ 77: 7000–7006. Article CAS PubMed PubMed Central Google Scholar * Rosset R, Julien J, Monier R . (1966). Ribonucleic acid composition of bacteria
as a function of growth rate. _J Mol Biol_ 18: 308–320. Article CAS PubMed Google Scholar * Schaechter M, Maaloe O, Kjeldgaard N . (1958). Dependency on medium and temperature of cell
size and chemical composition during balanced growth of Salmonella typhimurium. _Microbiology_ 19: 592. CAS Google Scholar * Schimel J, Balser T, Wallenstein M . (2007). Microbial
stress-response physiology and its implications for ecosystem function. _Ecology_ 88: 1386–1394. Article PubMed Google Scholar * Schippers A, Neretin LN, Kallmeyer J, Ferdelman TG, Cragg
BA, Parkes RJ _et al_ (2005). Prokaryotic cells of the deep sub-seafloor biosphere identified as living bacteria. _Nature_ 433: 861–864. Article CAS PubMed Google Scholar * Schmid M,
Schmitz-Esser S, Jetten M, Wagner M . (2001). 16S-23S rDNA intergenic spacer and 23S rDNA of anaerobic ammonium-oxidizing bacteria: implications for phylogeny and in situ detection. _Environ
Microbiol_ 3: 450–459. Article CAS PubMed Google Scholar * Simister R, Taylor MW, Tsai P, Fan L, Bruxner TJ, Crowe ML _et al_ (2012). Thermal stress responses in the bacterial biosphere
of the Great Barrier Reef sponge, Rhopaloeides odorabile. _Environ Microbiol_ 14: 3232–3246. Article CAS PubMed Google Scholar * Snyder L, Champness W . (2007) _Molecular Genetics of
Bacteria_ 3rd edn ASM Press: Washington, DC. Google Scholar * Sobek J, Charba J, Foust W . (1966). Endogenous metabolism of Azotobacter agilis. _J Bacteriol_ 92: 687–695. CAS PubMed
PubMed Central Google Scholar * Sukenik A, Kaplan-Levy RN, Welch JM, Post AF . (2012). Massive multiplication of genome and ribosomes in dormant cells (akinetes) of Aphanizomenon
ovalisporum (Cyanobacteria). _ISME J_ 6: 670–679. Article CAS PubMed Google Scholar * Tagkopoulos I, Liu Y-C, Tavazoie S . (2008). Predictive behavior within microbial genetic networks.
_Science_ 320: 1313–1317. Article CAS PubMed PubMed Central Google Scholar * Tissieres A, Watson JD . (1958). Ribonucleoprotein particles from Escherichia coli. _Nature_ 182: 778–780.
Article CAS PubMed Google Scholar * Tolker-Nielsen T, Larsen MH, Kyed H, Molin S . (1997). Effects of stress treatments on the detection of Salmonella typhimurium by in situ
hybridization. _Int J Food Microbiol_ 35: 251–258. Article CAS PubMed Google Scholar * van Bodegom P . (2007). Microbial maintenance: a critical review on its quantification. _Microb
Ecol_ 53: 513–523. Article PubMed PubMed Central Google Scholar * Verberkmoes NC, Denef VJ, Hettich RL, Banfield JF . (2009). Systems biology: functional analysis of natural microbial
consortia using community proteomics. _Nat Rev Microbiol_ 7: 196–205. Article CAS PubMed Google Scholar * Wade HE, Robinson HK . (1965). The distribution of ribosomal ribonucleic acids
among subcellular fractions from bacteria and the adverse effect of the membrane fraction on the stability of ribosomes. _Biochem J_ 96: 753–765. Article CAS PubMed PubMed Central Google
Scholar * Wagner R . (1994). The regulation of ribosomal RNA synthesis and bacterial cell growth. _Arch Microbiol_ 161: 100–109. Article CAS PubMed Google Scholar * Wilmes P, Bond PL .
(2004). The application of two-dimensional polyacrylamide gel electrophoresis and downstream analyses to a mixed community of prokaryotic microorganisms. _Environ Microbiol_ 6: 911–920.
Article CAS PubMed Google Scholar * Worden AZ, Binder BJ . (2003). Growth regulation of rRNA content in Prochlorococcus and Synechococcus (marine cyanobacteria) measured by whole-cell
hybridization of rRNA-targeted peptide nucleic acids. _J Phycol_ 39: 527–534. Article CAS Google Scholar * Wüst PK, Horn MA, Drake HL . (2011). Clostridiaceae and Enterobacteriaceae as
active fermenters in earthworm gut content. _ISME J_ 5: 92–106. Article PubMed Google Scholar * Yarwood S, Brewer E, Yarwood R, Lajtha K, Myrold D . (2013). Soil microbe active community
composition and capability of responding to litter addition after 12 yeears of no inputs. _Appl Environ Microbiol_ 79: 1385–1392. Article CAS PubMed PubMed Central Google Scholar *
Zinser ER, Lindell D, Johnson ZI, Futschik ME, Steglich C, Coleman ML _et al_ (2009). Choreography of the transcriptome, photophysiology, and cell cycle of a minimal photoautotroph,
Prochlorococcus. _PLoS ONE_ 4: e5135. Article PubMed PubMed Central Google Scholar Download references ACKNOWLEDGEMENTS We thank Jim Prosser, Josh Schimel, Eoin Brodie and Laurent
Philippot for constructive comments. SJB was supported by a National Science Foundation Graduate Research Fellowship. RLB was funded by the European Community’s Seventh Framework Programme
under grant agreement PIOF-GA-2008-219357. DOE Genomic Science Program grant (FOA DE-PS02-09ER09-25 award #0016377) to MKF. AUTHOR INFORMATION Author notes * Steven J Blazewicz Present
address: 4Current address: US Geological Survey, 345 Middlefield Road, MS 962, Menlo Park, CA 94025, USA., * Romain L Barnard Present address: 5Current address: INRA, UMR1347 Agroécologie,
17 rue Sully, BP 86510, Dijon, France., AUTHORS AND AFFILIATIONS * The Department of Environmental Science, Policy, and Management, University of California, Berkeley, CA, USA Steven J
Blazewicz, Romain L Barnard & Mary K Firestone * Department of Plant and Microbial Biology, University of California, Berkeley, CA, USA Rebecca A Daly * Ecology Department, Earth
Sciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA, USA Rebecca A Daly & Mary K Firestone Authors * Steven J Blazewicz View author publications You can also search
for this author inPubMed Google Scholar * Romain L Barnard View author publications You can also search for this author inPubMed Google Scholar * Rebecca A Daly View author publications You
can also search for this author inPubMed Google Scholar * Mary K Firestone View author publications You can also search for this author inPubMed Google Scholar CORRESPONDING AUTHOR
Correspondence to Steven J Blazewicz. ETHICS DECLARATIONS COMPETING INTERESTS The authors declare no conflict of interest. RIGHTS AND PERMISSIONS Reprints and permissions ABOUT THIS ARTICLE
CITE THIS ARTICLE Blazewicz, S., Barnard, R., Daly, R. _et al._ Evaluating rRNA as an indicator of microbial activity in environmental communities: limitations and uses. _ISME J_ 7,
2061–2068 (2013). https://doi.org/10.1038/ismej.2013.102 Download citation * Received: 31 October 2012 * Revised: 02 May 2013 * Accepted: 22 May 2013 * Published: 04 July 2013 * Issue Date:
November 2013 * DOI: https://doi.org/10.1038/ismej.2013.102 SHARE THIS ARTICLE Anyone you share the following link with will be able to read this content: Get shareable link Sorry, a
shareable link is not currently available for this article. Copy to clipboard Provided by the Springer Nature SharedIt content-sharing initiative KEYWORDS * community rRNA * microbial
activity * microbial growth * ribosomes * environmental samples * ecosystem processes
