Identification of the novel potential pathogen trueperella pecoris with interspecies significance by lamp diagnostics
- Select a language for the TTS:
- UK English Female
- UK English Male
- US English Female
- US English Male
- Australian Female
- Australian Male
- Language selected: (auto detect) - EN
Play all audios:

ABSTRACT _Trueperella pecoris_ was described as a new species of the genus _Trueperella_ in 2021 and might be pathogenic to various animal species. However, the lack of a suitable diagnostic
test system stands in the way of epidemiological surveys to clarify possible causalities. In this study, a Loop-mediated Isothermal Amplification (LAMP) assay was developed and validated
that was highly specific for _T_. _pecoris_. The assay provided an analytical sensitivity of 0.5 pg/25 µL and showed 100% inclusivity and exclusivity for 11 target and 33 non-target strains,
respectively. Three different DNA extraction methods were evaluated to select the most LAMP-compatible method for cell disruption in pure and complex samples. Using an on-site applicable
single-buffer DNA extraction with additional heating, the cell-based detection limit was 2.3 CFU/reaction. Finally, the LAMP assay was validated by means of artificially contaminated porcine
lung tissue samples in which minimal microbial loads between 6.54 and 8.37 × 103 CFU per swab sample were detectable. The LAMP assay established in this study represents a suitable
diagnostic procedure for identifying _T. pecoris_ in clinical specimens and will help to collect epidemiological data on the pathogenicity of this species. SIMILAR CONTENT BEING VIEWED BY
OTHERS ESTABLISHMENT AND VALIDATION OF A SIMPLE AND ACCURATE QPCR DETECTION METHOD FOR _HAEMOPHILUS PARASUIS_ Article Open access 10 March 2025 A NOVEL MULTIPLEX QPCR‑HRM ASSAY FOR THE
SIMULTANEOUS DETECTION OF FOUR ABORTIVE ZOONOTIC AGENTS IN CATTLE, SHEEP, AND GOATS Article Open access 28 July 2023 DESIGN AND VALIDATION OF _DOLOSIGRANULUM PIGRUM_ SPECIFIC PCR PRIMERS
USING THE BACTERIAL CORE GENOME Article Open access 14 April 2023 INTRODUCTION In 2021, _Trueperella_ (_T._) _pecoris_ was introduced as a novel species of the genus _Trueperella_ by
Schönecker et al.1. Strains were isolated from specimens of dairy cattle suffering from mastitis or abortion and of a dead pig showing fibrinous pleurisy and suppurative bronchopneumonia
during necropsy. In addition to this bacterium, other potential pathogens were also co-isolated, so that an etiological link could not be proven, but could not be excluded either. Based on
the anamnestic surveys and results of cultural examinations, the authors suspected at least a probable involvement of _T. pecoris_ in the pathogenesis of the mastitis cases described in that
study1. However, other species of the genus _Trueperella_, such as _T. abortisuis_ and _T. pyogenes_ have also been associated with pathological manifestations in pigs, including abortion
or multi-organ inflammation2,3. Recently, another _T. pecoris_ strain was isolated from the necrotic vestibulitis of a camel4. Apart from the previously described origins of bacterial
isolates and their phenotypic and genotypic characteristics, no information is available on this novel _Trueperella_ species. Its distribution and pathogenic significance remain unclear,
particularly as practicable, specific and reliable detection methods are lacking. Cultural examination followed by biochemical differentiation and sequencing of the 16S rRNA gene as
described by Schönecker et al.1 may already offer possibilities for identifying _T. pecoris_ but are very labour-intensive and time-consuming. To efficiently study the epidemiology of this
potential pathogen, rapid diagnostic methods suitable for screening large numbers of heterogeneous specimens are necessary. As detailed sequence information on various _T. pecoris_ strains
is available1, the development of molecular detection methods, for example based on Loop-mediated Isothermal Amplification (LAMP) should be considered. This technique was introduced in 20005
and since then has gained a lot of attention in the scientific community. Compared to alternative methods like PCR, LAMP enables rapid and continuous amplification at a constant
temperature, eliminating the need for an expensive and non-portable thermocycling device6. In addition, the method was shown to be highly specific and sensitive in detecting genomic targets
while being robust to inhibitory components in various samples7, 8. Reactions can be carried out at low cost, for example using a heating block9. For the subsequent detection of LAMP
products, various methods are available, which are often based on a visible color change or a generated turbidity10. In addition to low-cost options for performing this method, which can be
of great benefit in limited laboratory environments, there are also portable real-time fluorometers or turbidimeters available on the market11,12. These offer additional features like
real-time monitoring of the LAMP reaction or melting curve generation for evaluating the specificity of a LAMP product. The advantageous characteristics of the LAMP method have led to the
development of several assays intended as point-of-care diagnostic tools for a wide range of pathogens13. For the detection of microorganisms, LAMP is almost universally applicable as long
as a specific genomic target region is available. Therefore, the present study aimed to develop and validate an easy-to-perform LAMP assay for detecting _T. pecoris_ in clinical specimens.
Primarily, it should provide a diagnostic tool for further research into epidemiological aspects and pathogenic significance of this new species. Additionally, the assay should meet the
requirements for possible later use in patient diagnostics. RESULTS LAMP ASSAY SETUP A specific and conserved genomic target region within strain _T. pecoris_ 19OD0592 (Acc. No.
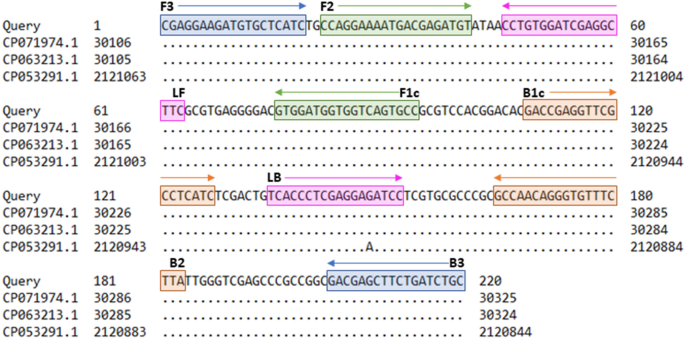
NZ_CP063212.1) was determined from 31,777 to 31,996 bp using the BLAST algorithm by the National Center for Biotechnology Information (NCBI) (https://blast.ncbi.nlm.nih.gov). Primer
sequences and positions are shown in Fig. 1. For setting up the _Trueperella pecoris_ (TP) LAMP assay, primer concentrations and reaction temperature were optimized. The concentrated TP
primer set resulted in more effective amplification and gave an advantage of about 5 min in detection time compared to the standard TP primer set (data not shown). Using the concentrated
primer set, the shortest detection times were measured at 65 °C (Fig. 2). Therefore, the concentrated primer set and the reaction temperature of 65 °C were selected as the final running
parameters. The positive control had a mean detection time of 9.26 ± 0.03 min with a LAMP product melting temperature of 90.7 ± 0.06 °C. In the following experiments, LAMP product melting
temperatures of 90.7 ± 1.0 °C were classified as specific for different isolates. ANALYTICAL SPECIFICITY The analytical specificity of the TP LAMP assay was evaluated by means of 44
bacterial strains including 11 T_. pecoris_ isolates of different origins (Table 1) and 33 non-target species. Inclusivity and exclusivity of the assay were both 100% with respect to the
isolates tested (Table 2). ANALYTICAL SENSITIVITY To determine the analytical sensitivity of the TP LAMP assay, ten-fold serially diluted DNA was tested. A minimum concentration of 0.5 pg
DNA/25 µL could be detected in all three repetitions. Decreasing DNA content in the reaction mixture was associated with a flattening of the amplification curves and a non-linear increase in
detection times (Fig. 3A + B). In contrast, a real-time PCR targeting the same amplicon as TP LAMP detected a minimum concentration of 50 fg DNA/25 µL in all three repetitions. CELL-BASED
DETECTION LIMITS Determining the cell-based detection limits of the TP LAMP assay was based on three DNA extraction methods. Using the DNeasy Blood & Tissue kit, up to 6.0 × 102 CFU/25
µL were detectable by TP LAMP in all three repetitions. This was 600 × less sensitive compared to the genomic target-identical real-time PCR, which was able to detect up to 1 CFU/20 µL. The
detection limit of the TP LAMP assay improved to 2.3 × 102 CFU/25 µL using single-buffer extraction and even reached 2.3 CFU/25 µL after an additional heating step was applied. DETECTION
LIMITS IN ARTIFICIALLY CONTAMINATED PORCINE LUNG TISSUE SAMPLES Artificially contaminated porcine lung tissue samples were tested to evaluate the application suitability of the TP LAMP assay
under the influence of sample material and background flora of the specimen type. The lung tissue was collected during dissections performed as a part of routine diagnostic procedures at
the Field Station for Epidemiology in Bakum (University of Veterinary Medicine Hannover). Detection limits for _T. pecoris_ strains DSM 111392T and 203/7 were determined and assessed using
three swab sampling adapted DNA extraction methods. Corresponding results in terms of pathogen count per swab are shown in Table 3. Using kit-extracted DNA, TP LAMP was 100- and 300-fold
less sensitive than real-time PCR for _T. pecoris_ strains DSM 111392T and 203/7, respectively. However, all results could be clearly interpreted and no false negatives occurred when samples
were tested with TP LAMP. In contrast, real-time PCR frequently produced false positive signals under the influence of the sample matrix. Melting curves of the LAMP products were always
uniform, whereas melting curves of the corresponding PCR products showed a wide variety of profiles compared to the positive control, with mostly diverging but also partly uniform melting
peaks. DISCUSSION _Trueperella pecoris_ is a novel bacterial species that appears to be related to several animal diseases, including mastitis in cattle, bronchopneumonia in pigs, and
impairments of the reproductive tract in ruminants1,4. In addition, the presence of _T. pecoris_ in the bovine mammary gland suggests that human infection through consumption of contaminated
milk should also be considered, as the human pathogenic properties of _Trueperella_ spp. are frequently described in the literature14, 15 However, the available data do not allow definitive
conclusions to be drawn as to whether _T. pecoris_ actually occurs as a disease-causing agent. In this study, a detection system was developed that enables specific diagnostics in clinical
specimens and thus helps to generate epidemiological data and to clarify the pathogenic significance of the bacterium. Primers of the TP LAMP assay were based on a gene region encoding an S9
family peptidase that has been shown to be unique in _T. pecoris_ by BLAST analysis. The functions of these enzymes have so far been best studied in humans and are associated with complex
immunological and hormonal metabolic processes16. It can be assumed that the targeted S9 family peptidase also plays a key role in the metabolism of _T. pecoris_ and that the coding gene is
constantly present and conserved in the bacterial genome. This was reflected in the high level of inclusivity of the TP LAMP assay for target species and exclusivity for non-target species.
Performances of TP LAMP and PCR targeting the same amplicon were compared in terms of analytical sensitivity. In the literature, there is true competition between the two amplification
techniques, often emphasizing that LAMP is superior to PCR10. However, this must be viewed in a differentiated manner. For example, LAMP usually achieves a higher analytical sensitivity only
when compared to conventional PCR17, 18. Considering results of real-time PCR, similar or even worse detection limits are frequently obtained19,20. Therefore, it was not unusual that the TP
LAMP assay presented in this study showed an analytical sensitivity 10 times lower than that of real-time PCR. The TP LAMP assay was able to compensate for this difference with other
advantageous properties and was superior to real-time PCR, particularly in the testing of complex sample matrices. When examining the cell-based detection limit, interestingly, there was a
significant loss in sensitivity when the kit-extracted DNA was tested. In contrast, the performance of the TP LAMP assay could be improved by LPTV buffer extraction with additional heating,
which was most likely due to a comparatively higher DNA yield. In addition, this DNA extraction method is characterized by its rapidity and its suitability for on-site use. However, no
washing steps are included, meaning that the inhibitory components are not removed. This regularly poses a problem for PCR diagnostics, as insufficiently processed DNA templates of low
quality can be associated with suppression of amplification and obtaining false negative results21. In contrast, LAMP is known for its ability to deal with potential inhibitors22, making the
method compatible with non-sophisticated, fast, and low-cost DNA preparation techniques. The TP LAMP assay also demonstrated the previously described properties when tested on porcine lung
tissue samples. This matrix was selected as it may represent a potential target organ for infection with _T. pecoris_1_._ Both tissue particles and blood adhered to the swab samples taken,
thus containing several inhibitory components like hemoglobin, lactoferrin, or immunoglobulin G23,24. Similar to the previous findings of this study, TP LAMP showed lower sensitivity for
kit-extracted DNA samples compared to real-time PCR, but compensated for this limitation when templates from LPTV buffer extraction with subsequent heating were used. However, differences of
TP LAMP and real-time PCR regarding sensitivity in complex samples can only be evaluated to a limited extent, as false positive reactions frequently occurred in real-time PCR examination.
In contrast, all TP LAMP reactions could be clearly interpreted and showed no false positive results, underpinning the robustness of this assay. Currently, no cultural or biomolecular
reference method is available for the diagnosis of _T. pecoris_, so that the identification of the bacterium is so far only possible via the non-directed collection of suspicious isolates
followed by laborious and time-consuming sequence analyses. Consequently, securing suitable sample material from infected animals during appropriate studies and subsequent testing by TP LAMP
would therefore be required to finally complete the validation of this assay. Since only the 11 _T. pecoris_ strains listed in Table 1 have been isolated to date, this process is expected
to take a longer period of time and could not be addressed in this study. No knowledge is yet available regarding the zoonotic potential of this species, but if transmission to humans occurs
through food-producing animals, validation of the TP LAMP assay should also be extended to meat and milk samples. In conclusion, the TP LAMP assay described in this study represents the
first available method for diagnosing _T. pecoris_ infections in complex sample materials. The fact that the assay detects the bacterium with high specificity and sensitivity in non-purified
templates reduces the costs, labor, and time for sample analysis, while at the same time ensuring high reliability of the results. These are suitable conditions for conducting large-scale
prevalence studies, since the TP LAMP assay can be used under both laboratory and field conditions. The data that can now be collected using TP LAMP diagnostics can be processed to draw
conclusions about the pathogenicity of _T. pecoris_ in humans and animals, which should be the subject of future studies. MATERIALS AND METHODS LAMP PRIMER The sequence of _T. pecoris_
strain 19OD0592 served as the basis for designing six specific LAMP primers using LAMP designer software version 1.15 (PREMIER Biosoft, Palo Alto, CA, USA). Sequence data was obtained from
the GenBank database (acc. No. NZ_CP063212.1) provided by NCBI (Bethesda, USA). For determining a species-specific sequence region, sections of the _T. pecoris_ genome were aligned
successively with other sequence data available at GenBank using the megablast algorithm of the integrated Basic Local Alignment Search Tool (BLAST) (NCBI). After importing the selected data
into the LAMP designer software, various LAMP primers were generated using default parameters and subjected to the integrated primer BLAST25. Subsequently, an appropriate primer set,
consisting of four basic and two loop primers, was selected based on the ranking by the software and according to optimal specificity properties (Fig. 1). Production of the LAMP primers was
contracted to Eurofins Genomics GmbH (Ebersberg, Germany). LAMP ASSAY OPTIMIZATION AND SETUP All LAMP reactions were carried out using the real-time fluorometer Genie II® by OptiGene Ltd.
(Horsham, UK). The device is equipped with a rechargeable battery and can be operated on site via touchscreen, which also allows direct result visualization. To determine the optimal
composition of the individual TP LAMP primers in a reaction mixture, two primer sets, consisting of a standard and a concentrated version, were prepared in accordance with the
recommendations of OptiGene Ltd. Using the standard TP primer set, concentrations of primers F3 and B3, primers FIP and BIP, and primers LF and LB in a reaction mixture were 0.2 µM, 0.8 µM,
and 0.4 µM, respectively. In contrast, concentrations of primers F3 and B3, primers FIP and BIP, and primers LF and LB in a reaction mixture were 0.2 µM, 2.0 µM, and 1.0 µM, respectively,
when the concentrated TP primer set version was used. Each reaction mixture with a total volume of 25 µL contained 15 µL GspSSD Isothermal Mastermix (ISO-001) (OptiGene Ltd.), 2.5 µL primer
mix, 2.5 µL nuclease-free water (NFW), and 5 µL DNA template. The optimal primer set was selected by detection times achieved for standardized DNA (0.1 ng/µL) from _T. pecoris_ reference
strain DSM 111392T and _T. pecoris_ field strain 203/7. Subsequently, the reaction temperature of the TP LAMP assay was optimized for the selected primer set by evaluating standardized DNA
(0.1 ng/µL) from _T. pecoris_ reference strain DSM 111392T using a temperature gradient between 62 and 69 °C on the Genie II® instrument. The temperature at which the lowest detection times
were measured was applied for subsequent LAMP tests. Optimization experiments were performed in triplicate and evaluated with the Genie Explorer version 2.0.7.11 application software
(OptiGene Ltd.) using default parameters. Each run included a positive and negative control reaction using _T. pecoris_ DSM 111392T DNA template (0.1 ng/µL) and NFW, respectively. REAL-TIME
PCR ASSAY A real-time PCR assay targeting the same amplicon as the TP LAMP assay was established to compare the performance of both amplification techniques. For this purpose, LAMP primers
TP-F3 and TP-B3 were used as forward and reverse primer in the real-time PCR assay, respectively. Each reaction had a total volume of 20 µL and contained 10 µL FastStart Essential DNA Green
Master (Roche), 1 µL of both TP-F3 and TP-B3 primer (10 µM), 3 µL H2O, and 5 µL DNA template. Runs started with initial denaturation at 95 °C for 10 min, followed by 45 cycles of three-step
amplification, and melting. Three-step amplification included denaturation at 95 °C for 10 s, annealing at 60 °C for 10 s and elongation at 72 °C for 10 s with ramp rates of 4.4, 2.2 and 4.4
°C. respectively. Fluorescence signals were measured after each elongation step. Melting curves were generated using a temperature–time profile of 95 °C for 10 s, 65 °C for 60 s and 97 °C
for 1 s. As with the TP LAMP assay, _T. pecoris_ DSM 111392T DNA template (0.1 ng/µL) and NFW were used as positive and negative control, respectively, in each run. DNA EXTRACTION FROM
BACTERIAL ISOLATES For DNA extraction, all bacteria were grown on Columbia sheep blood agar (Oxoid Deutschland GmbH, Wesel, Germany) in accordance with species-specific requirements.
_Escherichia coli_, _Enterococcus faecalis_ and _Enterococcus faecium_ isolates were incubated for 24 h at 37 °C under aerobic conditions. In contrast, a candle jar atmosphere was used to
cultivate _Trueperella_ spp. and _Arcanobacterium_ spp. for 24–48 h at 37 °C. For isolation of DNA, the DNeasy Blood & Tissue Kit (Qiagen GmbH, Hilden, Germany) was used with slight
modification. Instead of harvesting bacterial cells from liquid cultures by centrifuging, 5–10 colonies of Gram-positive or Gram-negative bacteria were picked from agar plates and directly
resuspended in enzymatic lysis buffer (20 mM Tris–HCl, pH 8.0; 2 mM sodium EDTA; 1.2% Triton® X-100; lysozyme, 20 mg/mL) or ATL buffer, respectively. The following steps for DNA extraction
were performed in accordance with the manufacturer's instructions. ANALYTICAL SPECIFICITY A total of 44 bacterial isolates were used to test the analytical specificity of the TP LAMP
assay (Table 2). Strains were derived from the inhouse culture collection of the Institute of Food Quality and Food Safety, University of Veterinary Medicine Hannover, Foundation (Hannover,
Germany), the German Collection of Microorganisms and Cell Cultures GmbH (Braunschweig, Germany), the National Collection of Type Cultures (Salisbury, UK), and from the RIPAC-LABOR GmbH
(Potsdam, Germany, ENOVAT European strain database). For inclusivity testing, previously described _T. pecoris_ strains originating from cattle and camel1,4 as well as further _T. pecoris_
isolates were considered (Table 1). Positive results were defined by the appearance of a fluorescence signal associated with a specific melting temperature. Strains used for exclusivity
testing were selected due to their close genetic relationship with _T. pecoris_ and their coexistence in the same environment. ANALYTICAL SENSITIVITY The analytical sensitivity of the TP
LAMP assay was determined by means of ten-fold serially diluted DNA from strain _T. pecoris_ DSM 111392T. AE buffer from the DNeasy Blood & Tissue Kit (Qiagen) was used as dilution
medium. DNA concentrations were adjusted to 1 ng/µL to 1 fg/µL and each template was tested in triplicate using TP LAMP. For comparison, each template was also examined three times by
real-time PCR. The analytical sensitivity was defined as the minimum DNA amount per reaction at which results were positive in all three replicates. DNA EXTRACTION FROM CELLS AND PORCINE
LUNG TISSUE Three different DNA extraction methods, including a kit-based method for obtaining highly purified DNA as well as a single-buffer extraction and a single-buffer extraction with
heating, both adapted for on-site application, were used to isolate DNA from _T. pecoris_ cell suspensions and porcine lung tissue swab samples. Kit-based extraction was performed using the
DNeasy Blood & Tissue kit (Qiagen) in accordance with the manufacturer’s recommendations. The required incubation steps at 37 °C and 56 °C lasted 1 h each. To compare the sensitivity of
LAMP and PCR, all kit-based isolates were tested with both methods. Applying the single buffer extraction method, pellets were suspended in 500 µL LPTV buffer (AmplexDiagnostics GmbH, Gars
am Inn, Germany), incubated for 5 min at room temperature and directly tested exclusively using TP LAMP. Subsequently, the same LPTV cell suspensions were subjected to heating at 100 °C for
10 min and tested again. CELL-BASED DETECTION LIMIT The cell-based detection limit of the TP LAMP assay was determined by ten-fold serially diluted cell-suspensions. For this purpose, strain
_T. pecoris_ DSM 111392T was grown for 48 h at 37 °C in a candle jar. Colony material was suspended in 5 mL NaCl solution (0.9%) until a turbidity of 2.0 McFarland units (MFU) was achieved.
Subsequently, the suspension was ten-fold serially diluted from 10–1 to 10–8 using 9-mL Maximum Recovery Diluent tubes (Oxoid Deutschland GmbH). DNA was extracted as previously described
from cell pellets obtained from 1 mL of each dilution level after centrifuging the 1-mL aliquots for 10 min at 7500 rpm. Supernatants were carefully discarded. Each isolation procedure was
repeated three times. The detection limit was defined as the minimum _T. pecoris_ cell count at which results were positive in all three replicates. ARTIFICIAL CONTAMINATION OF PORCINE LUNG
TISSUE SAMPLES For evaluating the suitability of the TP LAMP method under the influence of sample matrix and its background flora, swab samples were taken from porcine lung tissue and
artificially contaminated with _T. pecoris_. Strain _T. pecoris_ DSM 111392T and the camel-associated _T. pecori_s strain 203/7 were used for artificial contamination. For this purpose,
strains were grown for 48 h at 37 °C in a candle jar, and decimal dilution series were prepared as previously described. The surface of the lung of a clinically healthy pig as well as deeper
tissue layers opened by a sterile incision into the lung tissue were dabbed with a sterile cotton swab. Subsequently, the swab was transferred into a 1.5-mL reaction tube filled with 900 µL
MRD, repeatedly revolved, and finally squeezed out on the wall. For each strain, 10 tubes were prepared. From each dilution level of the two strain cell suspensions, 100 µL were taken and
added to the swab material containing MRD medium. One tube per strain remained non-inoculated and served as a negative extraction control. After centrifuging the tubes at 7500 rpm for 10
min, supernatants were discarded carefully. Pellets were used for DNA extraction as previously described. The experiment was repeated three times on three consecutive days. The detection
limit in porcine lung tissue samples was defined as the average _T. pecoris_ cell count per swab at which results were positive in all three replicates. DATA PROCESSING AND STATISTICAL
ANALYSES Data generated during the LAMP and PCR runs were analyzed using the Genie Explorer version 2.0.7.11 and Light Cycler® 96 software version 1.1.0.1320 application software provided by
the respective instrument manufacturers. Raw data processing and statistical analyses were performed with Microsoft Excel version 2303. DATA AVAILABILITY The datasets used and analysed
during the current study are available from the corresponding author on reasonable request. REFERENCES * Schönecker, L. _et al._ _Trueperella pecoris_ sp. nov. isolated from bovine and
porcine specimens. _Int. J. Syst. Evol. Microbiol._ https://doi.org/10.1099/ijsem.0.004848 (2021). Article PubMed Google Scholar * Jarosz, ŁS., Gradzki, Z. & Kalinowski, M.
_Trueperella pyogenes_ infections in swine: Clinical course and pathology. _Pol. J. Vet. Sci._ 17, 395–404. https://doi.org/10.2478/pjvs-2014-0055 (2014). Article CAS PubMed Google
Scholar * Alssahen, M. _et al._ Epidemiological analysis of _Trueperella abortisuis_ isolated from cases of pig abortion of a single farm. _Folia Microbiol._ 65, 491–496.
https://doi.org/10.1007/s12223-019-00753-9 (2020). Article CAS Google Scholar * Ahmed, M. F. E. _et al._ Phenotypic and genotypic approach to characterize a _Trueperella pecoris_ strain
isolated from necrotic vestibulitis of a camel (_Camelus dromedarius_). _Lett. Appl. Microbiol._ 75, 363–367. https://doi.org/10.1111/lam.13735 (2022). Article CAS PubMed Google Scholar
* Notomi, T. _et al._ Loop-mediated isothermal amplification of DNA. _Nucleic Acids Res._ 28, e63–e63. https://doi.org/10.1093/nar/28.12.e63 (2000). Article CAS PubMed PubMed Central
Google Scholar * Notomi, T., Mori, Y., Tomita, N. & Kanda, H. Loop-mediated isothermal amplification (LAMP): Principle, features, and future prospects. _J. Microbiol._ 53, 1–5.
https://doi.org/10.1007/s12275-015-4656-9 (2015). Article CAS PubMed Google Scholar * Wong, Y. P., Othman, S., Lau, Y. L., Radu, S. & Chee, H. Y. Loop-mediated isothermal
amplification (LAMP): A versatile technique for detection of micro-organisms. _J. Appl. Microbiol._ 124, 626–643. https://doi.org/10.1111/jam.13647 (2018). Article CAS PubMed Google
Scholar * Kaneko, H., Kawana, T., Fukushima, E. & Suzutani, T. Tolerance of loop-mediated isothermal amplification to a culture medium and biological substances. _J. Biochem. Biophys.
Methods_ 70, 499–501. https://doi.org/10.1016/j.jbbm.2006.08.008 (2007). Article CAS PubMed Google Scholar * Rodríguez-García, Á. _et al._ DNA extraction with DNAzol and LAMP, performed
in a heating block as a simple procedure for detection of _Mycobacterium tuberculosis_ in sputum specimens. _Methods Protoc._ https://doi.org/10.3390/mps1040037 (2018). Article PubMed
PubMed Central Google Scholar * Soroka, M., Wasowicz, B. & Rymaszewska, A. Loop-mediated isothermal amplification (LAMP): The better sibling of PCR?. _Cells_
https://doi.org/10.3390/cells10081931 (2021). Article PubMed PubMed Central Google Scholar * Kreitlow, A. _et al._ Establishment and validation of a loop-mediated isothermal
amplification (LAMP) assay targeting the ttrRSBCA locus for rapid detection of _Salmonella_ spp. in food. _Food Control_ 126, 107973. https://doi.org/10.1016/j.foodcont.2021.107973 (2021).
Article CAS Google Scholar * Zhang, Y. _et al._ Development of a _fimY_-based loop-mediated isothermal amplification assay for detection of _Salmonella_ in food. _Food Res. Int._ 45,
1011–1015. https://doi.org/10.1016/j.foodres.2011.02.015 (2012). Article CAS Google Scholar * Garg, N., Ahmad, F. J. & Kar, S. Recent advances in loop-mediated isothermal
amplification (LAMP) for rapid and efficient detection of pathogens. _Curr. Res. Microb. Sci._ 3, 100120. https://doi.org/10.1016/j.crmicr.2022.100120 (2022). Article CAS PubMed PubMed
Central Google Scholar * Deliwala, S., Beere, T., Samji, V., McDonald, P. J. & Bachuwa, G. When zoonotic organisms cross over-_Trueperella pyogenes_ endocarditis presenting as a septic
embolic stroke. _Cureus_ 12, e7740. https://doi.org/10.7759/cureus.7740 (2020). Article PubMed PubMed Central Google Scholar * Gilarranz, R., Chamizo, F., Horcajada, I. &
Bordes-Benítez, A. Prosthetic joint infection caused by _Trueperella bernardiae_. _J. Infect. Chemother._ 22, 642–644. https://doi.org/10.1016/j.jiac.2016.02.003 (2016). Article PubMed
Google Scholar * Page, M. J. & Di Cera, E. Serine peptidases: Classification, structure and function. _Cell Mol. Life Sci._ 65, 1220–1236. https://doi.org/10.1007/s00018-008-7565-9
(2008). Article CAS PubMed Google Scholar * Priya, G. B. _et al._ Development and evaluation of isothermal amplification assay for the rapid and sensitive detection of _Clostridium
perfringens_ from chevon. _Anaerobe_ 54, 178–187. https://doi.org/10.1016/j.anaerobe.2018.09.005 (2018). Article CAS PubMed Google Scholar * Hara-Kudo, Y., Yoshino, M., Kojima, T. &
Ikedo, M. Loop-mediated isothermal amplification for the rapid detection of _Salmonella_. _FEMS Microbiol. Lett._ 253, 155–161. https://doi.org/10.1016/j.femsle.2005.09.032 (2005). Article
CAS PubMed Google Scholar * Anupama, K. P., Nayak, A., Karunasagar, I., Karunasagar, I. & Maiti, B. Evaluation of loop-mediated isothermal amplification assay along with conventional
and real-time PCR assay for sensitive detection of pathogenic _Vibrio parahaemolyticus_ from seafood sample without enrichment. _Mol. Biol. Rep._ 48, 1009–1016.
https://doi.org/10.1007/s11033-020-06116-9 (2021). Article CAS PubMed PubMed Central Google Scholar * Cao, X. _et al._ Detection of viable but nonculturable _Vibrio parahaemolyticus_ in
shrimp samples using improved real-time PCR and real-time LAMP methods. _Food Control_ 103, 145–152. https://doi.org/10.1016/j.foodcont.2019.04.003 (2019). Article CAS Google Scholar *
Schrader, C., Schielke, A., Ellerbroek, L. & Johne, R. PCR inhibitors—Occurrence, properties and removal. _J. Appl. Microbiol._ 113, 1014–1026.
https://doi.org/10.1111/j.1365-2672.2012.05384.x (2012). Article CAS PubMed Google Scholar * Yang, Q., Wang, F., Prinyawiwatkul, W. & Ge, B. Robustness of _Salmonella_ loop-mediated
isothermal amplification assays for food applications. _J. Appl. Microbiol._ 116, 81–88. https://doi.org/10.1111/jam.12340 (2014). Article CAS PubMed Google Scholar * Al-Soud, W. A.
& Rådström, P. Purification and characterization of PCR-inhibitory components in blood cells. _J. Clin. Microbiol._ 39, 485–493. https://doi.org/10.1128/JCM.39.2.485-493.2001 (2001).
Article CAS PubMed PubMed Central Google Scholar * Al-Soud, W. A., Jönsson, L. J. & Rådström, P. Identification and characterization of immunoglobulin G in blood as a major
inhibitor of diagnostic PCR. _J. Clin. Microbiol._ 38, 345–350. https://doi.org/10.1128/JCM.38.1.345-350.2000 (2000). Article CAS PubMed PubMed Central Google Scholar * Altschul, S. F.
_et al._ Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. _Nucleic Acids Res._ 25, 3389–3402. https://doi.org/10.1093/nar/25.17.3389 (1997). Article CAS
PubMed PubMed Central Google Scholar Download references ACKNOWLEDGEMENTS The authors would like to thank Silke Ortaeri and Anke Bertling for their excellent technical assistance. Thanks
also go to Manfred Merle for editing the illustrations for publication. FUNDING This Open Access publication was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research
Foundation) - 491094227 “Open Access Publication Funding” and the University of Veterinary Medicine Hannover, Foundation. Open Access funding enabled and organized by Projekt DEAL. AUTHOR
INFORMATION AUTHORS AND AFFILIATIONS * Institute of Food Quality and Food Safety, University of Veterinary Medicine Hannover, Bischofsholer Damm 15, 30173, Hannover, Germany Antonia
Kreitlow, Madeleine Plötz & Amir Abdulmawjood * Faculty of Veterinary Medicine, Universitas Wijaya Kusuma Surabaya, Jl. Dukuh Kupang XXV No.54, Dukuh Kupang, Surabaya, 60225, Indonesia
Siti Gusti Ningrum * Institute of Hygiene and Infectious Diseases of Animals, Justus Liebig University Giessen, Frankfurter Str. 87-89, 35392, Giessen, Germany Christoph Lämmler &
Christiane Hoffmann * RIPAC-LABOR GmbH, Am Mühlenberg 11, 14476, Potsdam, Germany Marcel Erhard Authors * Antonia Kreitlow View author publications You can also search for this author
inPubMed Google Scholar * Siti Gusti Ningrum View author publications You can also search for this author inPubMed Google Scholar * Christoph Lämmler View author publications You can also
search for this author inPubMed Google Scholar * Marcel Erhard View author publications You can also search for this author inPubMed Google Scholar * Christiane Hoffmann View author
publications You can also search for this author inPubMed Google Scholar * Madeleine Plötz View author publications You can also search for this author inPubMed Google Scholar * Amir
Abdulmawjood View author publications You can also search for this author inPubMed Google Scholar CONTRIBUTIONS A.K., C.L., and A.A. were responsible for the conceptualization of the
project. A.K. designed the experiments and wrote the original draft of the manuscript. A.K. and S.G.N. performed experiments. A.K., S.G.N, and A.A. analyzed the data. M.E. and C.H. provided
_T. pecoris_ strains. M.P. supervised the project. All authors revised and approved the final version of the paper. CORRESPONDING AUTHOR Correspondence to Amir Abdulmawjood. ETHICS
DECLARATIONS COMPETING INTERESTS The authors declare no competing interests. ADDITIONAL INFORMATION PUBLISHER'S NOTE Springer Nature remains neutral with regard to jurisdictional claims
in published maps and institutional affiliations. RIGHTS AND PERMISSIONS OPEN ACCESS This article is licensed under a Creative Commons Attribution 4.0 International License, which permits
use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the
Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless
indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory
regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit
http://creativecommons.org/licenses/by/4.0/. Reprints and permissions ABOUT THIS ARTICLE CITE THIS ARTICLE Kreitlow, A., Ningrum, S.G., Lämmler, C. _et al._ Identification of the novel
potential pathogen _Trueperella pecoris_ with interspecies significance by LAMP diagnostics. _Sci Rep_ 13, 14005 (2023). https://doi.org/10.1038/s41598-023-40787-1 Download citation *
Received: 02 June 2023 * Accepted: 16 August 2023 * Published: 27 August 2023 * DOI: https://doi.org/10.1038/s41598-023-40787-1 SHARE THIS ARTICLE Anyone you share the following link with
will be able to read this content: Get shareable link Sorry, a shareable link is not currently available for this article. Copy to clipboard Provided by the Springer Nature SharedIt
content-sharing initiative
